
Sergio E. Pasteris
INSIBIO-UNT-CONICET
Title: Clonal analysis and virulence-associated traits of native Escherichia coli from urethra of gilts and natural/artificial pregnant sows.
Submitted Date: 2018-01-30
Biography
Sergio E. Pasteris has his expertise in the study of lactic acid bacteria (LAB) metabolism as well as in the isolation and evaluation of its beneficial properties to design probiotics for amphibians´ culture. Taken into account the development of resistant bacteria, some LAB strains represent an alternative instead chemotherapeutics to prevent epizootics in bullfrog systems breeding. Since bullfrog is a carrier of the etiological agent of chytridiomycosis, probiotics by using native LAB from bullfrog skin are being developed to be applied during the ex situ breeding of endangered amphibian species.
Abstract
Statements of the Problem: The reproductive performance of sows is a key factor in the herd´s productivity. Urinary tract infections (UTI) are a common problem in females, causing repeat breeding with a delayed return to estrus, which reduces the animal´s welfare and the litter performance; Escherichia coli being associated to these infections. Diverse studies described a unique microbiota in the UT in bitch. Others authors concluded that the composition of the UT bacterial communities could have an important role in the health condition of the host.\r\n\r\nMethodology: We performed the isolation and clonal association (rep-PCR, Box and Eric primers) of E. coli from the urethral microbiota of: healthy gilts-HG (n=9) and pregnant sows by natural breading-NB (n=11) or artificial insemination-AI (n=11). Also, 12 virulence factors relevant for pyelonephritic strains were evaluated by PCR: hlyA, cnf, ibeA, iutA, kpsMT II, FimH, papC, sfa/focD, afa/draBC, traT, agn43, csgA. \r\n\r\nFindings: Cultures revealed a slightly minor count (CFU/mL) for AI (3.7±0.59) group compared to HG (4.2±0.24) and NB (4.3±0.44). However, there were no differences for E. coli isolation (CFU/mL): 1.45±1.36, 2.87±1.53 and 2.61±1.84, for AI, NB and HG, respectively. The clonal analysis with both, Box or Eric primers, revealed a high similarity (>90%) between E. coli isolates from different animal groups. Positive reaction was found for: FimH (76%), agn43 (92%), traT (32%) and csgA (72%), these last ones showed a differential prevalence and were associated with E. coli from NB sows. \r\n\r\nConclusion & Significance: These results indicate that the management conditions could affect the characteristics of the urethral microbiota in sows and, therefore, the risk for urinary tract diseases. \r\n\r\nFigure: Conditions in the reproductive management (AI or NM) affect the characteristics of the urinary tract microbiota by favoring the prevalence of pathogenic microorganisms and thus, decreasing the reproductive performance in sows. UTI, Urinary Tract Infections; AI, Artificial Insemination; NM, Natural Malting (Breading)\r\n
Sergio E Pasteris
INSIBIO-CONICET
Title: Probiotics for amphibians: advances in the selection of lactic acid bacteria for chytridiomycosis control
Submitted Date: 2018-01-30
Biography
Sergio E Pasteris has his expertise in the study of lactic acid bacteria (LAB) metabolism as well as in the isolation and evaluation of its beneficial properties to design probiotics for amphibian culture. Taken into account the development of resistant bacteria, some LAB strains represent an alternative instead chemotherapeutics to prevent epizootics in bullfrog systems breeding. Since bullfrog is a carrier of the etiological agent of chytridiomycosis, probiotics by using native LAB from bullfrog skin are being developed to be applied during the ex situ breeding of endangered amphibian species.\r\n \r\n
Abstract
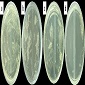
Statement of the Problem: Chytridiomycosis is an amphibian skin disease caused by Batrachochytrium dendrobatidis (Bd) that produces extinction around the world. Some amphibian-skin bacteria have been proposed as probiotic for chytridiomycosis control but they excluded the lactic acid bacteria (LAB) group. Here in, we advanced in the selection of indigenous LAB from bullfrog (considered as a Bd carrier) skin to design probiotic for application during the ex situ breeding of endangered amphibians. \r\n\r\nMethodology: To determine the anti-Bd activity, co-culture assays between Bd strains (CFLT 159 from Brazil; AVS4 and AVS7 from Chile) and potentially LAB isolates were performed. Isolates previously shown exopolysaccharide (EPS) synthesis and/or auto aggregation (AA) ability were evaluated for biofilm formation by using polyestyrene plates. Compatibility assays were performed to evaluate the possibility to formulate a mixed probiotic. \r\n\r\nFindings: From 62 potentially LAB, 48 isolates had any anti-Bd activity. The 16s RNA sequence analysis allowed obtaining 97-99% of identity that matches with Enterococcus and Lactobacillus. Thus, Enterococcus sp. 90, 564, 747, 762; Lactobacillus sp. 10, 529, and Enterococcus gallinarum CRL 1826 (previously characterized) inhibited all the Bd strains. Three LAB isolates exhibited low biofilm formation, while E. gallinarum showed moderate production. This ability was not always associated with AA or EPS synthesis. The compatibility assays indicated that the LAB isolates could be included in mixed probiotic with the exception of Enterococcus sp. 742 that was inhibited by E. gallinarum. \r\n\r\nConclusion & Significance: E. gallinarum CRL 1826 resulted the best strain for a probiotic since it has many beneficial properties: anti-Bd activity, AA, EPS synthesis, biofilm formation, medium hydrophobicity and GRAS properties according to in vitro and in vivo tests (3-7). However, Enterococcus sp. 747 would maximize some probiotic properties of the CRL strain; therefore, a mixed probiotic can be proposed.\r\n\r\nFigure: Anti-Bd activity of LAB. a- Enterococcus sp. 81 against Bd AVS7: negative inhibition; b- Enterococcus sp. 564 against Bd CFLT 159: low inhibition; c-Enterococcus sp. 747 against Bd AVS4: medium inhibition; d- E. gallinarum CRL 1826 against Bd AVS7: strong inhibition.\r\n

